Inputs of Research and development activities in ECOMORE 2 project
 Project coordinator: Elodie CALVEZ
Project coordinator: Elodie CALVEZ
Member staffs: Souksakhone VIENGPHOUTHONG, Phaithong BOUNMANY
Ecomore2 project in Lao PDR aimed at gathering multiple sources of data for the development of an outbreak simulator for the dengue virus. Viral genetic information and the level of herd immunity of the Lao population against the dengue virus were considered as key information for the accuracy of the simulator. Accordingly, specific tools were designed for the rapid analysis of large series of human samples.
From a genetic point of view, the four dengue virus serotypes are subdivided into genotypes based on genetic distances and phylogenetic relationships. The full-length sequence of the envelop gene is used as the gold standard to determine the dengue virus genotypes.
In 2013, the arbovirus surveillance network detected the emergence of DENV-4 in the Vientiane Capital. In 2014-2015, it spreads at the country level and DENV-4 became predominant in 2017-2018. Meanwhile, since 2012, DENV-2 was regularly detected at low levels in Lao PDR. Its epidemiologic profile changed in 2018.
Indeed, in October 2018, DENV-2 burden progressively increased, and DENV-2 became predominant in the Vientiane Capital in 2019 and at the country level, being responsible for a huge epidemic as mentioned previously. Regarding DENV-1, this serotype circulates in Lao PDR for years but has not been associated with a recent outbreak, though, since 2016, it represents about 22% of the samples serotyped by the arbovirus surveillance network. DENV3 has not been detected over the last years.
To determine the DENV-4 polymorphism observed in the Vientiane Capital and at the country level, we sequenced the full-length envelope gene of DENV-4 samples isolated during the major DENV-4 outbreak that occurred between 2014 and 2018. Obtained sequences were submitted to phylogenetic analyses. The results demonstrated the introduction of genotype II in 2014 (2 imported cases). However, since 2015, the only genotype I was detected on the autochthonous cases (Figure 5).
Regarding DENV-2 polymorphism, a panel of 49 DENV- 2 isolates, gathering both autochthonous and imported cases collected in six Lao provinces between 2015 to 2019, was analyzed. The molecular epidemiology revealed the co-circulation of two DENV-2 genotypes (Cosmopolitan and Asian genotypes) and of different lineages within each genotype over the last five years. Sequence comparison suggested multiple events of the introduction of DENV-2 strains from neighboring countries (Figure 6).

Figure 5. Maximum likelihood phylogenetic tree of DENV-4 sequences from Lao PDR. The tree was constructed based on the envelope gene (1,485 bp). Only bootstrap values > 70 are shown. Scale bar indicates the nucleotide substitution per site. The Lao strains sequenced in this study are indicated in red. The Lao strain previously described is indicated in blue. Black triangles highlight isolates obtained from fatal cases (Calvez E. et al., Pathogens 2020;9(9):728).
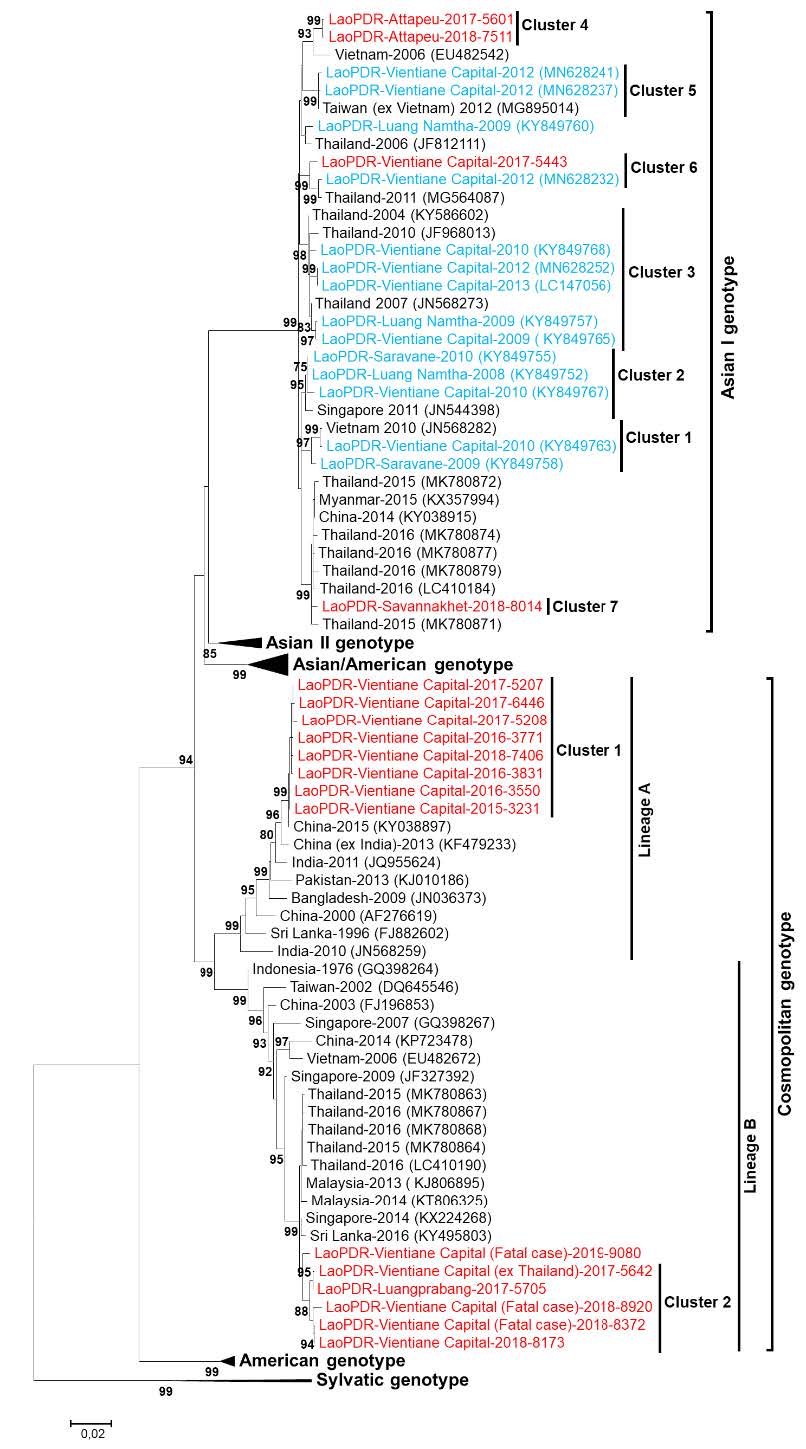
Figure 6. Maximum-likelihood phylogenetic tree of DENV-2 sequences from Lao PDR. The tree was constructed on a 1485 bp segment of the envelope gene. Only bootstrap values >70 are shown. Scale bar indicates nucleotide substitution per site. The Lao PDR strains analyzed in this study are indicated in red. The Lao PDR strains previously described are in blue. Blue clusters 1 to 3 were previously described by Castonguay-Vanier J. et al. PNTD 2018;12(1):e0006203. (Calvez E. et al., PlosOne 2020;15(8):e0237384).
For the last five years, DENV-1 has been circulating in Lao PDR, representing about 20% of the samples serotyped by the arbovirus surveillance network, but no recent outbreak was associated with this serotype. To better understand its circulation in Lao PDR and at the regional scale, we analyzed the envelope sequence of strains isolated since 2015. The preliminary results demonstrated the co-circulation of several clusters of DENV-1 Genotype I as regularly described in Southeast Asia.










